Genoinseq: Services
Services NGS
Genoinseq provides high-throughput sequencing and bioinformatics data analysis services since 2007. Our accumulated experience and specialized multidisciplinary team favor our focus on delivering personalized solutions and reliable support. We assist our clients in the experimental design of their projects, sequence with the most efficient and adequate approach for optimal results and help transform sequencing data into biological or clinical answers through a comprehensive set of bioinformatics tools.
Genoinseq additionally provides the technological platform to the University of Coimbra's Center for Neuroscience and Cell Biology (CNC) and the Center for Innovation in Biomedicine and Biotechnology (CIBB).
Genoinseq is a member of GenomePT (RNIE). ![]()
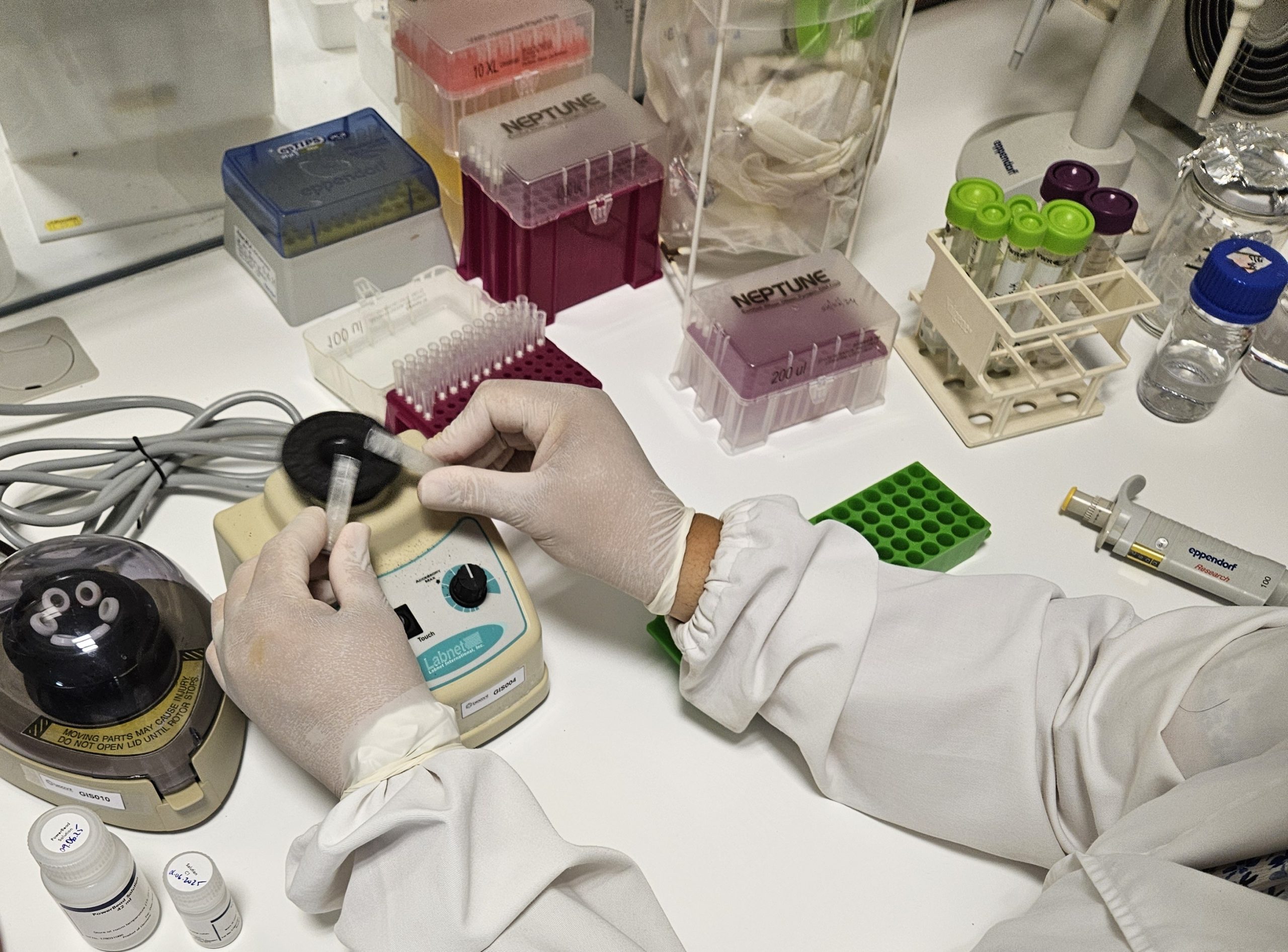

We currently have three next-generation sequencers (NGS) suited for different applications: Ion Proton™, from Thermo Fisher Scientific, MiSeq® and NextSeq® from Illumina®.
Ion Proton™, a semiconductor sequencer, generates more than 15 gigabases per sequencing run, with 60-80 million high-quality reads of 200 bp (maximum length).
MiSeq®, a highly flexible sequencer with short run times, provides up to 2X300 bp sequences and typical outputs of more than 15 gigabases with 22-25 million high-quality reads, per run.
NextSeq®, a high-throughput benchtop sequencer, has an output up to 100 Gb of 2X150 bp reads with 400 million high-quality reads, per run.
Applications
Genoinseq provides complete sequencing, genotyping and bioinformatics applications to meet your needs. We assist in the experimental design of your unique projects, sequence with the most efficient and adequate approach for optimal results and transform sequencing raw data into biological or clinical answers through a comprehensive set of bioinformatics tools.
Our vast experience in high throughput sequencing comes from working with a wide range of organisms such as humans, animals, plants, and microorganisms in different applications.
We are determinedly committed to quality, through the continuous improvement of our internal procedures to guarantee high-quality results.
Appropriate for genome sequencing of new or not yet sequenced organisms. Genoinseq can sequence genomes of bacteria, plasmids, fosmids, viruses, or fungi on the MiSeq® or NextSeq®.
Our service also includes bioinformatics data analysis. After sequencing and quality control, the reads are assembled into contigs, and the genes predicted and annotated with PGP V2 (Prokaryote Gene Prediction*). Outputs include QC, raw data (FASTQ), assembly metrics, contigs (FASTA) and gff or GenBank annotation files. Assistance in data deposition in public databases is included.
*Egas et al., 2014, doi: 10.4056/sigs.5661021
Genome resequencing provides a catalog of sequence variation to study diversity in individuals. Depending on the genome size or the number of samples, genomes can be re-sequenced in the MiSeq® or NextSeq® platforms according to genome size or sample number.
After sequencing and quality control, reads are mapped against the reference genome to identify small variants (SNPs, small insertions, and deletions) and structural variations (larger insertions and deletions). Variants are annotated for inferred effects (e.g. sequence conservation, type of variant, effect prediction on gene product or regulatory region) if annotation is available. Outputs include QC, raw data (FASTQ), alignment files (e.g. BAM files) and variant annotation files (e.g. VCF format).
Exome sequencing consists on sequencing all the exons in the human genome to identify genetic variation responsible for Mendelian diseases (85% of disease-causing mutations are in exons) or associated with complex diseases or health-related traits. Genoinseq sequences exomes on the NextSeq® and Ion Proton™.
For exome analysis, our laboratory developed ExomeLoupe, a pipeline to detect, annotate and visualize exon variants. The user can filter variants by gene, genotype, structural annotation, variant effect, known polymorphisms, population frequencies or phenotypes (HPO)
With targeted gene sequencing panels, one can select the genes or gene regions relevant to address a biological or clinical question, such as the genes associated or suspect to be associated with a particular disease or phenotype. The main advantage over exome or genome sequencing is the high sequencing depth.
Gene panels can be selected based on known gene-disease associations (ready-to-use panels) or can be designed to include genes or genomic regions of interest (custom panels). Genoinseq sequences gene panels on the MiSeq®, NextSeq® and Ion ProtonTM platforms.
This application measures gene expression through deep-sequencing to study the extent and complexity of a transcriptome. RNA-Seq allows for the discovery and annotation of complete transcripts or characterizes alternative splicing and polyadenylation. Depending on the genome size or the number of samples, RNA samples can be sequenced in the NextSeq® or MiSeq® platforms. De novo whole transcriptome sequencing is also available.
In RNA-Seq results analysis sequencing reads are mapped against an annotated reference genome or transcriptome. After normalization, reads are counted and analyzed with specific statistical software. Outputs include QC and raw data (FASTQ). Differential gene expression and annotation are accessed through a user-friendly web platform to visualize and explore results.
Targeted gene expression is an alternative approach to whole transcriptome sequencing to study selected genes. Solutions comprise predesigned panels or customizable panels which can include genes of interest. Genoinseq sequences gene expression panels on the MiSeq®, NextSeq® or Ion ProtonTM platforms.
This application is appropriate for small RNA discovery or study of expression patterns. These 10-40-nucleotide sequences play a major role in post-transcriptional regulation of gene expression. Small RNA includes small interfering RNAs (siRNAs), microRNAs (miRNAs) or Piwi-associated RNAs (piRNAs).
Once sequenced on the MiSeq® or NextSeq® platforms, reads are mapped against the reference genome of the organism under study or small RNA databases. Differential expression is calculated upon read count, normalization, and statistical analysis. Outputs include QC, raw data (FASTQ), differential expression and annotation information.
High-throughput sequencing is broadly used to characterize the composition and diversity of microbial communities from natural environments. This amplicon-based sequencing involves the amplification of a specific gene, usually the 16S ribosomal RNA gene for bacteria or the Internal Transcribed Spacer (ITS2) region for fungi. The approach is also successful for the identification of members of eukaryotic communities through sequencing of the 18S rRNA or the mitochondrial Cytochrome Oxidase I (COI) genes.
Upon sequencing on the MiSeq® platform, reads are quality filtered, merged and matched to reference taxa, or grouped into clusters at a fixed level of sequence identity named Operational Taxonomic Units (OTU) and assigned to reference taxa based on sequence homology. Outputs include QC, raw data (FASTQ), OTU taxonomical identification and respective abundance per sample, and alpha-diversity and beta-diversity when applicable.
This shotgun sequencing method is suitable for studying the taxonomic and functional profile of microbial communities. This application involves DNA fragmentation and small fragment deep sequencing. Genoinseq can sequence metagenomes from low to medium diversity microbial communities on the MiSeq® and NextSeq®.
Reconstruction of the taxonomical and functional metagenome profile involves a direct comparison of reads, or contigs, to a reference catalog of microbial genes or genomes. Functional profiling can also be inferred by comparison to databases of proteins or protein domains such as KEGG orthology, Clusters of Orthologous Groups (COGs), Swiss-Prot, Pfam or InterPro.
Microsatellite markers are an important tool for ecological and evolutionary research. For many non-model organisms where such markers are still unavailable or for which there is no genome sequence, random partial genome sequencing can help discover new microsatellite markers. Genoinseq can sequence on the MiSeq® or NextSeq®. Outputs include QC, raw data (FASTQ) or microsatellite identification with specialized software.
Genoinseq can assist in finding the application best suited for your needs. Please contact our services if you need further assistance. We can work with you to establish a sequencing approach or an adequate bioinformatics pipeline to best answer your biological/clinical questions.
Products
Genoinseq considers strategic to develop new products to improve services and applications to clients.
GenoinVar
GenoinVar is an end-to-end solution from blood sample to exome sequencing and variant analysis to accelerate the discovery of causal genetic variants. This solution was developed in the framework of the In2Genome project, a multidisciplinary project to integrate whole exome sequencing in clinical practice routine.
GenoinVar involves whole exome sequencing of extracted DNA on the Illumina NextSeq platform and variant analysis thought ExomeLoupe(link), an intuitive and user-friendly Windows software for variant prioritization and interpretation.
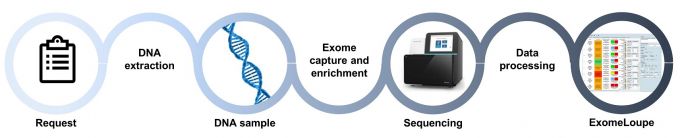
Main features:
- Done under a certified environment (ISO 9001).
- Great efficiency in the internal pipeline.
- Reduction in time to results.
- Security of the genetic data in all steps.
- Support from a multidisciplinary team of experts.
Under this end-to-end solution we tested molecular diagnosed patients within the In2Genome project and successfully identified the previously reported variants. GenoinVar represents a true end solution for identification of causal variants in clinical or research contexts.
ExomeLoupe
ExomeLoupe is an intuitive and user-friendly Windows platform to help researchers identify causal variants in a secure environment. The platform was developed in the framework of the In2Genome project, a multidisciplinary project to integrate whole exome sequencing in clinical practice routine.
ExomeLoupe displays the variants and most relevant annotations according to queries defined by the user in an informative and concise manner together with a colour code making the analysis more intuitive. The user can select the variants of interest by gene, HPO term or disease; genomic position; clinical significance based on ClinVar; population frequency; and/or predictive effect according to different tools. ExomeLoupe works on encrypted data, also enabling file storage and sharing in a protected manner according to the new European General Data Protection Regulation.


